 When the updated sequence file is successfully read, the Update Sequence dialog opens and displays four sections: Alignment, Sequence Update, Existing Features, and Options.Īlignment describes and displays the existing sequence and updated sequence, showing differences in various colors. 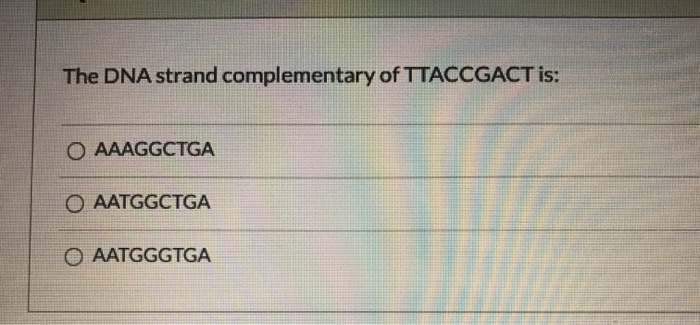
Choose the correct FASTA or ASN.1 file from the list of available files. When Update Sequence is chosen, your computer’s open/choose file dialog is opened. The dialog updates a single sequence at a time. Update Sequence allows the user to upload a new sequence to replace an existing sequence and make appropriate changes to existing features. The Cancel button will exit the dialog and undo any edits that have been made by the dialog. Note that when making changes in the Sequence Editing dialog, it is necessary to use the Commit button to apply changes to the data before adding new features with the Features menu or retranslating coding regions with the Retranslate button. When features are displayed, their locations can be adjusted by dragging the endpoints of their intervals. The user can edit the sequence directly by clicking on the sequence and typing characters or using backspace or delete. Selecting a sequence range is useful because the Annotate menu can be used to create a feature for the selected location. The “ Select:” text box enables a user to select a range of sequence without using the mouse to drag the cursor. This control will also allow the user to search for sequence characters – for example, searching for “atg” will move the cursor to the next instance of this codon. This is useful for navigating to a specific position without scrolling. The “ Go to:” text box enables the user to select the position of the cursor. The Edit->Find menu item launches a dialog that allows the user to search for sequence characters in the nucleotide sequence, the reverse complement, or in translated frames. The protein currently associated with the coding region is also displayed, and the user can choose Mismatch to highlight the positions on the protein sequence that do not match the calculated translation. For coding regions, the On-the-fly option shows the protein translation calculated using the sequence underlying the coding region and the frame of the coding region. The user could also choose to display features as labeled lines below the sequence or hide them. For example, the user could choose to view reading frames or display the sequence complement below the sequence. The Show menu controls the information displayed. The Edit Sequence dialog is a useful tool for viewing the sequence and features and editing the sequence content and feature locations. At any point to end the process without making changes simply click the cancel button. The new sequences will be added to the end of the file. When everything is correct the Problems column (The last column) will be empty. Simply type in the new ID in the New Sequence ID box and click the recheck problems button. 
In the new Sequence ID display of the example, it indicates that the problem is a duplicate ID. The bottom displays the new Sequence IDs. The top portion of the window displays all Sequence IDs from the existing sequence. If the imported file has the same sequence identifier as one of the existing files, an error will occur: The browse function can be used to find files to open or the files can simply be dropped in and recently used files will appear in the Recently used Files box for easy use. This can be changed to NCBI ASN.1 files or fasta alignment files. The default file format setting is fasta sequence files. Use Add Sequences to add the plasmids to the project. For example, the nuclear sequences of a bacterial genome already exist in a project, but the plasmid sequences weren’t included. The Add Sequences function can be used to add new sequences to an existing project already in progress. 
Expand Known Gaps to Include Flanking Ns.Reverse Complement Sequences by Sequence ID.Question 3 combines these two steps without any hints on the orientation, i.e., it just gives and expects the sequences without explicitly giving the 5' and 3' ends. Question 2 adds the second step, reversing the sequence to give the proper 5'-3' orientation. Question 1 simulates the first step, finding the complementary sequence. 
Give the DNA sequence that will pair with the following stretches of DNA. This usually involves reversing the sequence after writing it complementary to the one you are given. Remember, when writing complementary DNA sequences, you need to write the sequence in the 5' Because of the nature of complementary base pairing, if you know the sequence of one strand of DNA, you can predict the sequence of the strand that will pair with, or "complement" it. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
